Menu
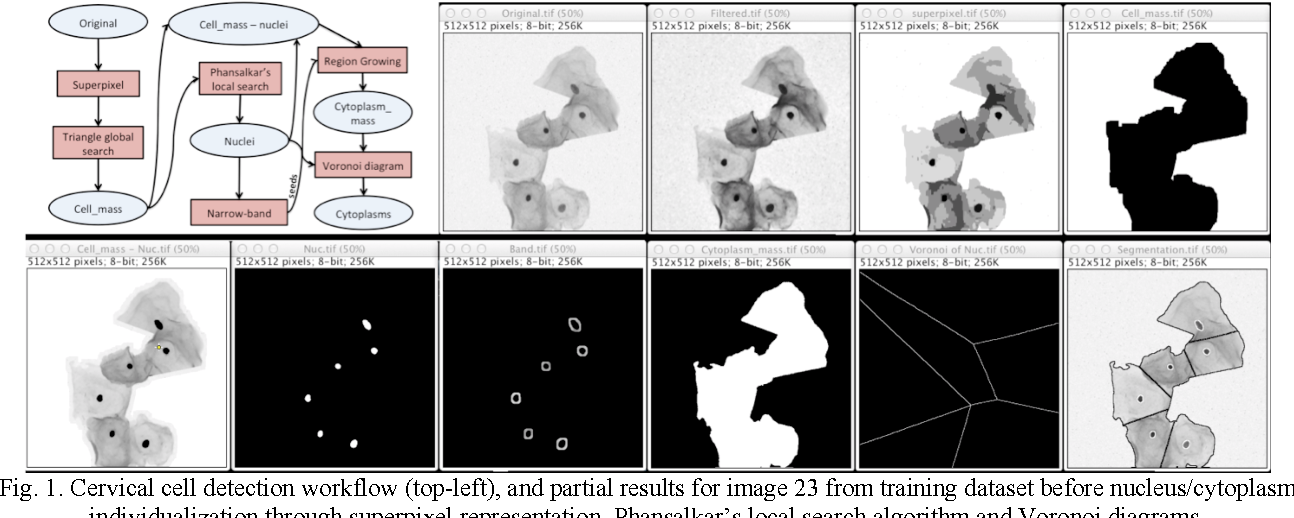
The segmentation step consists of classifying the small cropped images from the new set of test images using a pixel-wise sliding window as input to the CNN. For each test image, the CNN model with a sliding window technique creates a probability map, where each pixel is assigned to a probability of being an abnormal cell or not. Jan 5, 2015 - Segmentation of subcellular compartments combining superpixel representation with Voronoi diagrams. United States: N.
A Software Based Novel Approach: Integrated Segmentation & Nuclei Extraction of Overlapped Cervical Cell in High Resolution MRI ImagesInternational Journal of Engineering Trends and Technology (IJETT)© 2017 by IJETT JournalVolume-49 Number-4Year of Publication: 2017Authors: Setu Garg, Shabana Urooj, RituVijay10.5381/IJETT-V49P233Setu Garg, Shabana Urooj, RituVijay 'A Software Based Novel Approach: Integrated Segmentation & Nuclei Extraction of Overlapped Cervical Cell in High Resolution MRI Images', International Journal of Engineering Trends and Technology (IJETT), V49(4),206-211 July 2017. Published by seventh sense research groupAbstractIn this paper, we have proposed an algorithm to detect and locate the nuclei from cell masses and then segmentation ofoverlapped cervical in MRI images. Detection of cancerous area with fine boundaries in overlapped cell clusters is an important butchallenging task.In this proposed approach, multiple cell masses are detected from MRI images of overlapped cervical cells by generating superpixels and then multiple thresholding is performed. From cell masses, nuclei of multiple cells are detected and located by applying clustering thresholding. Finally, an integrated active contour algorithm is applied on cell nuclei for segmentation.References1 Sironi S, Zanello A, Rodighiero MG, Vanzulli A, Taccagni GL, Belloni C, Del Maschio A. Invasive carcinoma of the cervix uteri (Stage IB-IIB).
Comparison of CT and MR for the assessment of the parametrium. Radiol Med.81(5):671-7, May 1991.3 Kim SH, Choi BI, Han JK,Kim HD, Lee HP, Kang SB, Lee JY, Han MC. Preoperative staging of uterine cervical carcinoma: Comparison of CT and MRI in 99 patients. J Comput AssistTomogr, 17(4):633-40, Jul-Aug1993.4 Subak LE, Hricak H, Powell CB, Azizi E, Stern JL. Cervical Carcinoma: Computed tomography and Magnetic resonance imaging for preoperative staging. Obstetrics & Gynecology.86(1):43-50, July 1995.5 Parkin DM, Bray FI, &Devesa SS.
Cancer burden in the year 2000. The global picture.
European journal of cancer, 37(8):S4-66, Oct 2001.6 Chan TF, and Vese LA. Active contours without edges. IEEE transactions on Image Processing.
10(2):266-277, Feb 2001.7 Rousson M, and Paragios N. Shape Priors for Level Set Representations. European Conference on Computer Vision.
LNCS 2351: 78-92, May 2002.8 Jeronimo J, and Schiffman M. A Tool for Collection of Region Based Data from Uterine Cervix Images for Correlation of Visual and Clinical Variables Related to Cervical Neoplasia. Computer based medical systems. 17th IEEE Symposium.
558-562, June 2004.9 Matas J, Chum O, Urban M,Pajdla T. Robust wide-baseline stereo from maximally stable extremal regions. Image and Vision Computing. 22(10): 761-767, 2004.10 Arora S, Acharya J, Verma A, and PrasantaPanigrahi K.
Multilevel thresholding for image segmentation through a fast statistical recursive algorithm. Elsevier, Pattern Recognition Letters, 29: 119-125, 2008.11 Radau P, Lu Y, Connelly K, Paul G, Dick AJ, and Wright GA. Evaluation framework for algorithms segmenting short axis cardiac MRI. The MIDAS Journal – Cardiac MR Left Ventricle Segmentation Challenge. 2009.12 Achanta R, Shaji A, Smith K, Lucchi A, Fua P, and Susstrunk S. Slic superpixels compared to state-of-the-art superpixel methods. IEEE Transactions on Pattern analysis Machine Intelligence.
34(11): 2274-2282, Nov2012.13 Nosrati MS, and Hamarneh G. A variational approach for overlapping cell segmentation. Overlapping Cervical Cytology Image Segmentation Challenge – ISBI 2014.
2014.14 Ushizima DM, Bianchi AGC, and Carneiro CM. Segmentation of subcellular compartments combining superpixel representation with voronoi diagrams. Overlapping Cervical Cytology Image Segmentation Challenge – ISBI 2014, 2014.15 Phoulady HA, Goldgof DB, Hall LO, and Mouton PR. An approach for overlapping cell segmentationin multi-layer cervical cell volumes.
Overlapping Cervical Cytology Image Segmentation Challenge – ISBI 2015, 2015.16 Ramalho GLB, Ferreira DS, Bianchi AGC, Carneiro CM, Medeiros FNS, and Ushizima DM. Cell reconstruction under voronoi and enclosing ellipses from 3d microscopy. Overlapping Cervical Cytology Image Segmentation Challenge – ISBI 2015, 2015.17 Lu Z, carneiro G, and Bradley A. An improved joint optimization of multiple level set functions for the segmentation of overlapping cervical cells.IEEE Transactions on Image Processing. 24(4): 1261-1272, April 2015.18 PatilShweta R, and Patil VS.
Similarity measurement using shape feature for image retrieval. Jan 2016.19 R. Pradeep Kumar Reddy, Dr. C.Nagaraju,and I. Rajasekhar Reddy. Canny scale edge detection.IJETT- ICGTETM-N3.
Jan 2016.KeywordsCell Mass, Clustering Thresholding, Integrated Active Contour Method Multiple Thresholding, and Superpixel Generation.
Authors:;; Publication Date: 2015-01-05 Research Org.: Lawrence Berkeley National Lab. (LBNL), Berkeley, CA (United States) Sponsoring Org.: USDOE Office of Science (SC) OSTI Identifier: 1167389 Report Number(s): LBNL-6892E DOE Contract Number: DE-AC02-05CH11231 Resource Type: Conference Resource Relation: Conference: International Symposium on Biomedical Imaging, Apr 2014 Country of Publication: United States Language: English Subject: 97 MATHEMATICS AND COMPUTING; cervical cell quantification; Voronoi; segmentation.
Three-dimensional mammary cell culture models offer new opportunities for the development of computational techniques for segmentation, localization, and multicellular organization. Under normal conditions, these assays form a symmetrical, hollow structure, which is necessary for their functional operation. Often, the nuclear compartments are labeled, which provides context for quantitative protein localization or colony structure through fluorescent microscopy. These colonies are first delineated from the background using the level set method. Within each colony, nuclear regions are then bounded by their center of mass through iterative radial voting, and a local neighborhood for each nucleus is established through Voronoi tessellation. Finally, the level set method is applied again within each Voronoi region to delineate the nuclear compartment. The paper concludes with the application of the proposed method to a set of experimental data demonstrating a stable solution when iterative radial voting and level set methods are used synergistically.
Furthermore, segmented colonies are characterized for architectural changes as a result of ionizing radiation. Three dimensional cell culture models offer new opportunities for development of computational techniques for segmentation and localization.
These assays have a unique signature of a clump of cells that correspond to a functioning colony. Often the nuclear compartment is labeled and then imaged with fluorescent microscopy to provide context for protein localization. These colonies are first delineated from background using the level set method. Within each colony, nuclear regions are then bounded by their center of mass through radial voting, and a local neighborhood for each nucleus is established through Voronoi tessellation.
Finally, the level set method is applied again within each Voronoi region to delineate the nuclear compartment. The paper concludes with the application of the proposed method to a dataset of experimental data demonstrating a stable solution when iterative radial voting and level set methods are used synergistically. Three dimensional cell culture assays have emerged as the basis of an improved model system for evaluating therapeutic agents. Molecular probes, and exogenous stimuli.
However, there is a gap in robust computational techniques for segmentation of image data that are collected through confocal or deconvolution microscopy. The main issue is the volume of data, overlapping subcellular compartments, and variation in scale and size of subcompartments of interest. A geometric technique has been developed to bound the solution of the problem by first localizing centers of mass for each cell and then partitioning clump of cells along minimal intersecting surfaces. An approximate solution to the center of mass is realized through iterative spatial voting, which is tolerant to variation in shape morphologies and overlapping compartments and is shown to have an excellent noise immunity. These centers of mass are then used to partition a clump of cells along minimal intersecting surfaces that are estimated by Radon transform. Examples on real data and performance of the system over a large population of data are evaluated. Although proposed strategies have been developed and tested on data collected through fluorescence microscopy, they are applicable to other problems in low level vision and medical imaging.
Cell membrane proteins play an important role in tissue architecture and cell-cell communication. We hypothesize that segmentation and multidimensional characterization of the distribution of cell membrane proteins, on a cell-by-cell basis, enable improved classification of treatment groups and identify important characteristics that can otherwise be hidden. We have developed a series of computational steps to (i) delineate cell membrane protein signals and associate them with a specific nucleus; (ii) compute a coupled representation of the multiplexed DNA content with membrane proteins; (iii) rank computed features associated with such a multidimensional representation; (iv) visualize selected features for comparative evaluation through heatmaps; and (v) discriminate between treatment groups in an optimal fashion. The novelty of our method is in the segmentation of the membrane signal and the multidimensional representation of phenotypic signature on a cell-by-cell basis. To test the utility of this method, the proposed computational steps were applied to images of cells that have been irradiated with different radiation qualities in the presence and absence of other small molecules. These samples are labeled for their DNA content and E-cadherin membrane proteins. We demonstrate that multidimensional representations of cell-by-cell phenotypes improve predictive and visualization capabilities among different treatment groups, and identify hidden variables.
Biological membranes reflect a fascinating example of a separation system that is multifunctional, tunable, precise, and efficient. Biomimetic membranes, which mimic the architecture of cellular membranes, have the potential to deliver significant improvements in specificity and permeability. In this work, a fully synthetic biomimetic membrane is introduced that incorporates ultra-efficient 1.5 nm diameter carbon nanotube porin (CNTPs) channels in a block-copolymer matrix.
It is demonstrated that CNTPs maintain high proton and water permeability in these membranes. CNTPs can also mimic the behavior of biological gap junctions by forming bridges between vesicular compartments that allow transport of small molecules.
